About Me
I am an Associate Professor of Computational Chemistry in the Department of Chemistry at San José State University. I earned a Ph.D. in Theoretical Physical Chemistry from the University of California, Irvine, where my dissertation research was focused on the development and application of enhanced sampling algorithms for the computation of conformational kinetics in macromolecules from molecular dynamics simulations. My M.S. in Medicinal Chemistry, B.S., and B.A. were all completed at Villanova University. I was also a Data Science Fellow at UC Irvine and worked as a postdoctoral researcher at the California Institute for Telecommunications and Information Technology (Calit2) at UC Irvine, resulting in broad training in a variety of computational techniques including a variety of machine learning methodologies, network statistical modeling, and creating custom data visualizations. My research has been published in a variety of journals across disciplines, including the Journal of Chemical Information and Modeling, Nature Structural and Molecular Biology, the Journal of Chemical Physics, the Journal of Physical Chemistry, Biomolecules, the Journal of Chemical Education, and the SIAM Journal on Applied Mathematics. I have taught a wide variety of subjects ranging from physical chemistry, to programming in Python, to graduate computational chemistry, to science communication, and have even taught guitar professionally. In addition to my research and teaching experience, I have both past industry experience in a management role at the Merck Pharmaceutical Testing Lab as well as current industry experience as a technical consultant in the AI tools development space, where I leverage more than a decade of programming experience (Python, R, C/C++, Mathematica). I have also accrued over 1 million total views of my educational content across multiple YouTube channels, as I am passionate about science education outreach for communicating the broader impacts of science to scientists, people outside of the scientific community, and future scientists alike.
About the Grazioli Research Group
In a nutshell, we develop computational methods that use molecular simulations and machine learning to study configurational dynamics in biomolecules, molecular self-assembly, and chemical reaction dynamics. Our research interests are highly interdisciplinary, drawing from computational/theoretical chemistry and biophysics, physical chemistry, chemical physics, data science and structural biology. I aim to build on my research experience in enhanced sampling methodologies for molecular simulations, machine learning, and coarse-grained modeling toward building computational discovery methods for the molecular sciences. Highly motivated students of all levels and backgrounds, who are interested in learning to use computers to study chemistry and biophysics, are encouraged to reach out to me via my SJSU email. The work that we do is fundamentally interdisciplinary, so my group is a great fit for students from a wide variety of majors (e.g. chemistry, physics, computer science, engineering, bioinformatics, biology, etc.). No programming is experience required, only a willingness to learn, a willingness to work with others, and tenacity! For more information, please watch the video below, as well as the FAQ (frequently asked questions) video on my contact page:
Recent Publications
In addition to learning about my research interests here, please visit my Google Scholar profile for a list of my publications.
Machine Learning Applications in Chemistry and Biophysics
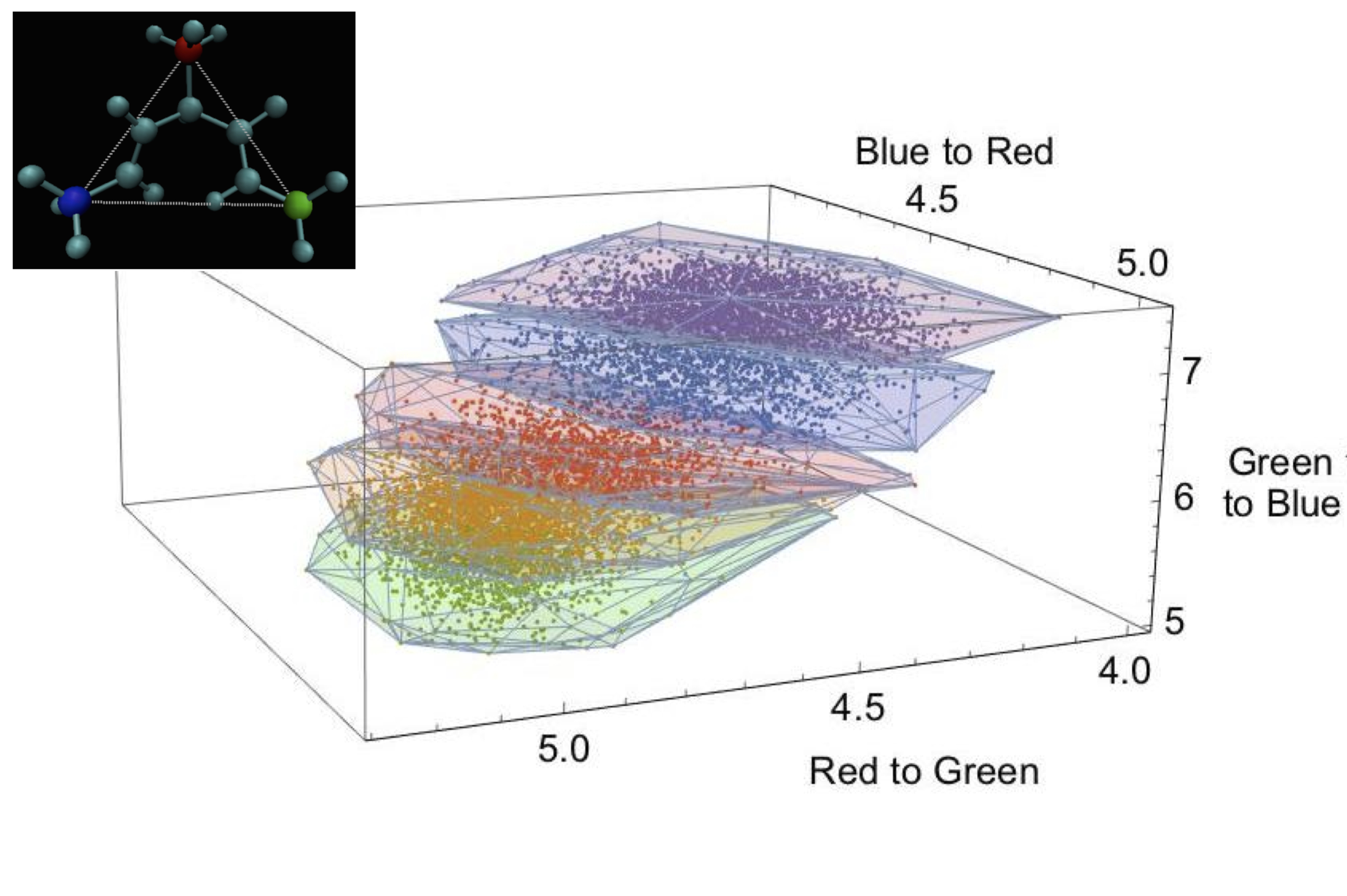
One area of focus for my research is the development of machine learning-based methods for both guiding and interpreting the results of molecular simulations. I am also interested in developing machine learning-based methodologies for interpreting experimental data.
Coarse-grained Modeling
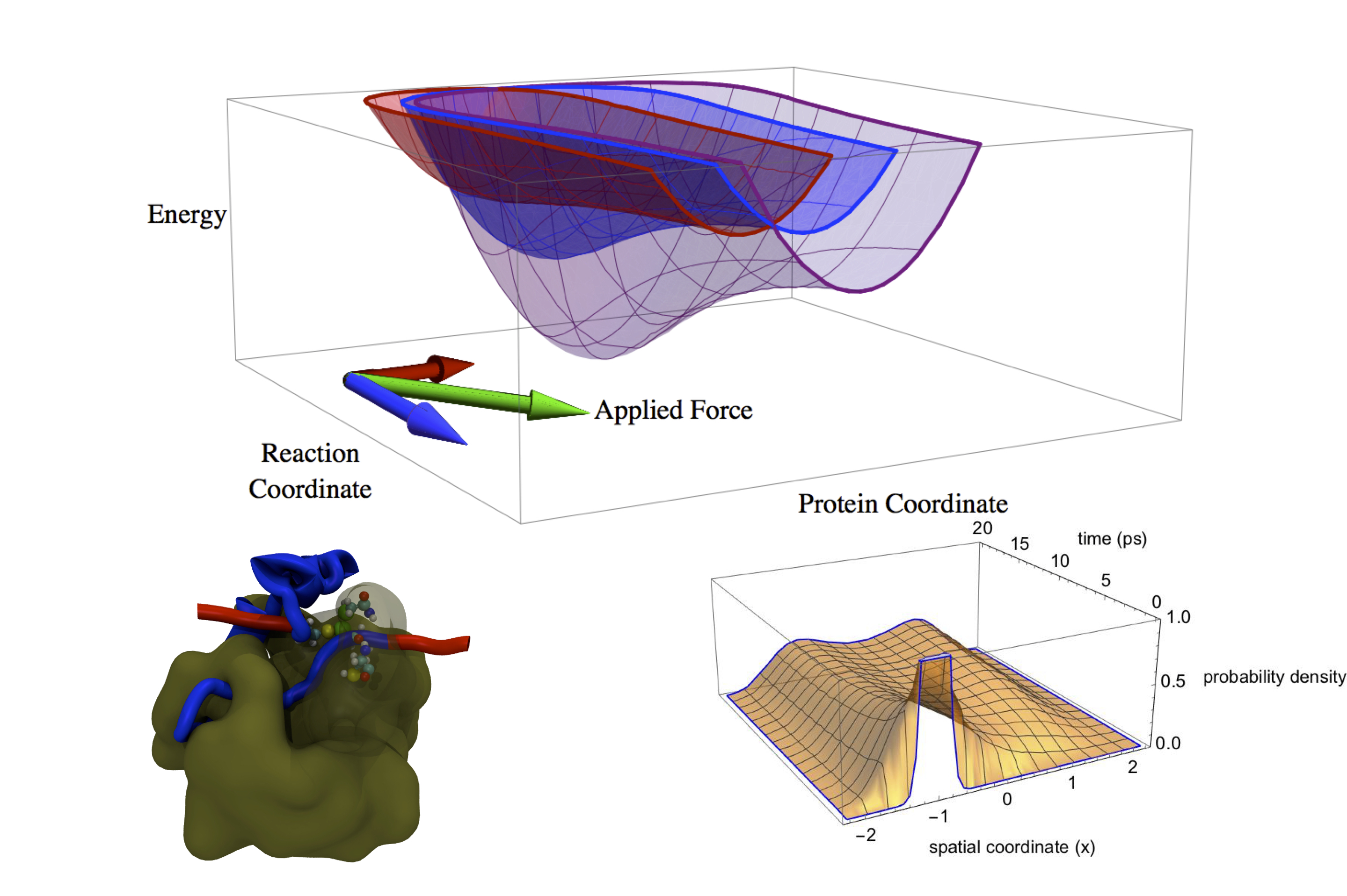
Another research focus of mine is on the development of coarse-grained modeling techniques for molecular systems whose complexity make atomistic simulations intractable on the time scales of interest. I have built coarse-grained models of phenomena such as protein aggregation for amyloid fibril formation and force-modulated catalytic activity in enzymes.
Molecular Dynamics Methodology
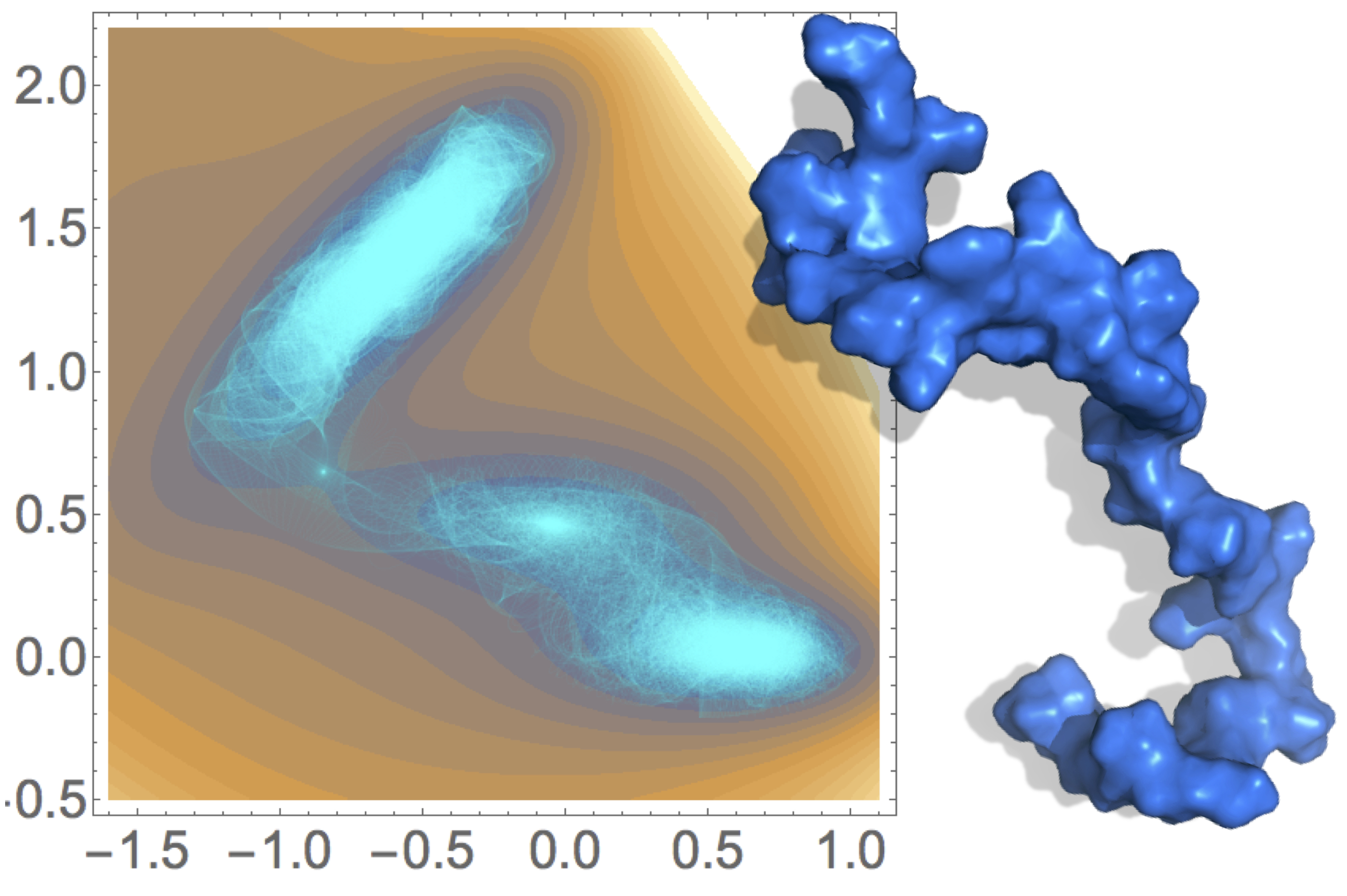
Much of my work in this area has been devoted toward creating enhanced sampling algorithms for calculating time-dependent properties from molecular simulations. I have developed multiple enhanced sampling methodologies for molecular dynamics simulations of complex systems, such as proteins and nucleic acids, as well as for exploring ab initio dynamics simulations of smaller molecules.